Projects
To understand co-evolution of leukemic and immune cell populations, we are interested in (i) lineage-tracing techniques that leverage natural genetic barcodes to map out clonal evolution following (immuno)-therapeutic bottlenecks such as allogeneic stem cell transplantation, and (ii) multi-omics approaches to define the phenotypes and specificities of therapeutically relevant T cell populations.
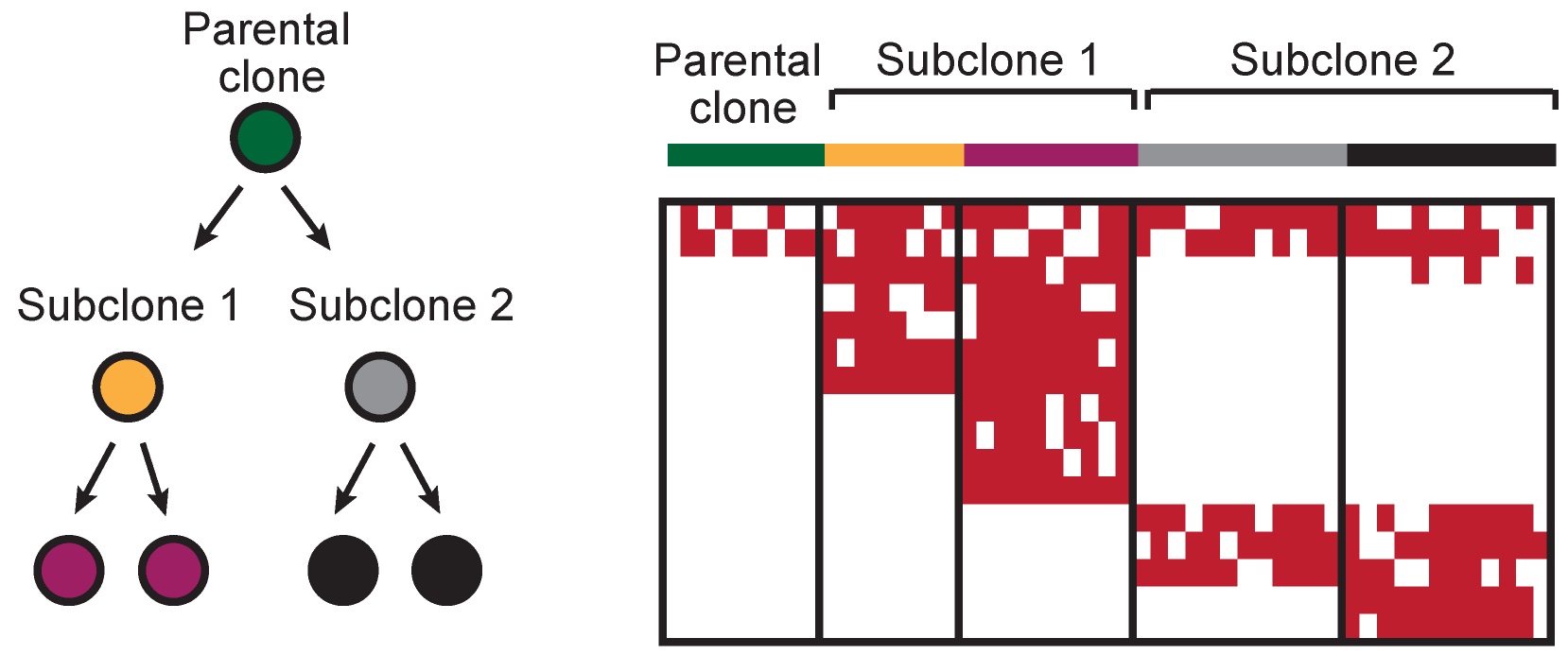
Lineage-tracing using natural genetic barcodes
The availability of approaches to resolve genetic and phenotypic information within the same cell opens up opportunities to better understand how immune resistance mechanisms develop that provide selective advantage and fuel clonal evolution.
We use multi-omics single cell data to dissect longitudinal changes in the cell state of leukemic clones defined by natural genetic barcodes and to gain novel insights into mechanisms that underpin relapse after immunotherapy.
We are also excited about the possibility to integrate multiple genetic barcodes (mitochondrial and somatic nuclear DNA mutations, copy number changes, expressed germline single nucleotide polymorphisms) to track rare malignant cells and to distinguish donor- from recipient-derived cells in the post-transplant context.
Leukemia-specific T cell responses
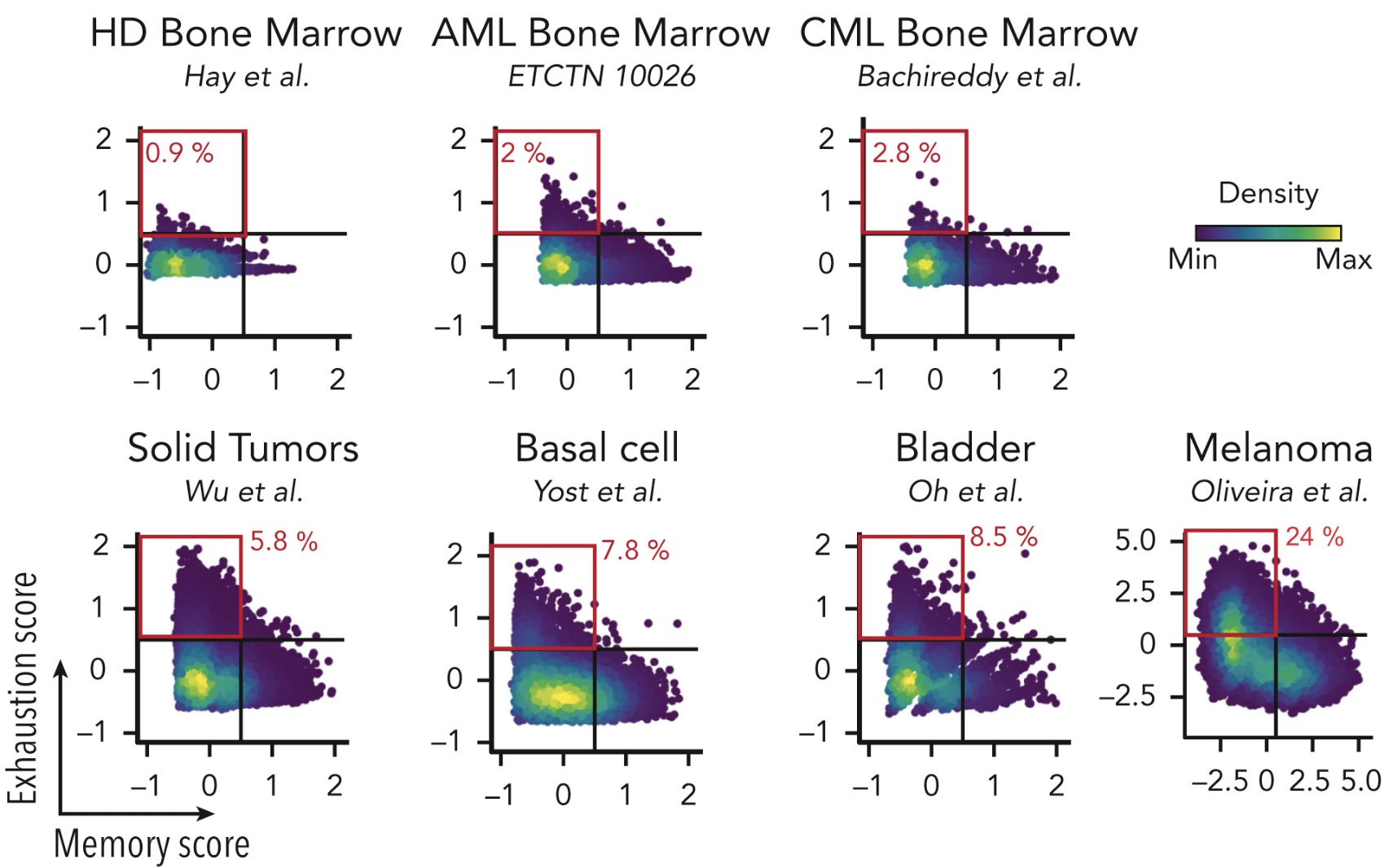
While immune checkpoint blockade and adaptive T cell therapy are transformative immunotherapies in solid tumors, they have so far seen much less success in blood malignancies. This likely relates to differences in mutational burden, the frequency of tumor-specific T cells and their phenotypes between hematologic and solid malignancies. A deeper understanding of the phenotypes and dynamics of leukemia-specific T cells is needed to advance T cell therapies for blood cancer.
We use single cell and spatial sequencing approaches to characterize and track therapeutically relevant T cells, for example in the context of allogeneic stem cell transplantation. We are also excited about understanding the phenotypes of T cells in extramedullary manifestations of AML, given observed sensitivity of leukemia cutis to CTLA-4 blockade.